How to Build a RAG Pipeline with UltraRAG v2 and Milvus
Retrieval-Augmented Generation (RAG) has evolved far beyond the simple “retrieve then generate” pattern. Modern systems now behave more like full reasoning engines, combining adaptive retrieval, multi-step planning, and dynamic decision-making. But this progress comes with two major challenges: high engineering cost and growing system complexity. Reproducing existing methods often requires rebuilding intricate pipelines, while experimenting with new ideas demands significant orchestration work.
UltraRAG v2 tackles these pain points directly. Developed by THUNLP, NEUIR, OpenBMB, and AI9stars, it’s the first RAG framework built on the Model Context Protocol (MCP). Instead of hand-writing complex logic, researchers can declare sequences, loops, and branching behavior in simple YAML files, enabling fast, low-code construction of multi-stage RAG systems.
Even with a framework like UltraRAG, a RAG system still needs a strong retrieval layer. This is where a vector database helps. Milvus, an open-source vector database, stores embeddings, builds indexes, and performs fast similarity search on large datasets. In an UltraRAG pipeline, Milvus retrieves relevant information for the model. Together, UltraRAG and Milvus make it easier to build RAG systems that are both flexible and efficient.
In this post, we’ll show how to integrate Milvus with UltraRAG v2 and build a complete RAG pipeline.
UltraRAG v2 Architecture at a Glance
In different RAG systems, core functions such as retrieval, generation, and evaluation are similar but implemented in different ways, making components difficult to reuse or combine. The MCP addresses this problem by standardizing how LLMs communicate with external tools through a simple Client–Server architecture.
Inspired by this, UltraRAG v2 is built around three core ideas:
- Modular encapsulation: UltraRAG v2 packages key RAG capabilities into standalone MCP Servers with unified Tool interfaces. This creates a clean, modular structure where you can focus on reasoning logic rather than backend wiring. New components can be added, replaced, or upgraded like plugins—no core-code edits required.
- YAML configuration: Complex, multi-step RAG pipelines are hard to debug. UltraRAG v2 makes them transparent by moving all control logic into YAML. Sequences, loops, and conditional branches are defined in a declarative manner, and every step’s inputs and outputs are clearly traceable. This greatly simplifies debugging and accelerates workflow iteration.
- Lightweight workflow orchestration: A built-in MCP Client executes pipeline, keeping workflow behavior fully decoupled from underlying implementations. While traditional RAG systems often require editing core code to add new features, UltraRAG v2 adopts a microservice-like model where new modules can be deployed independently, just like installing a plugin.
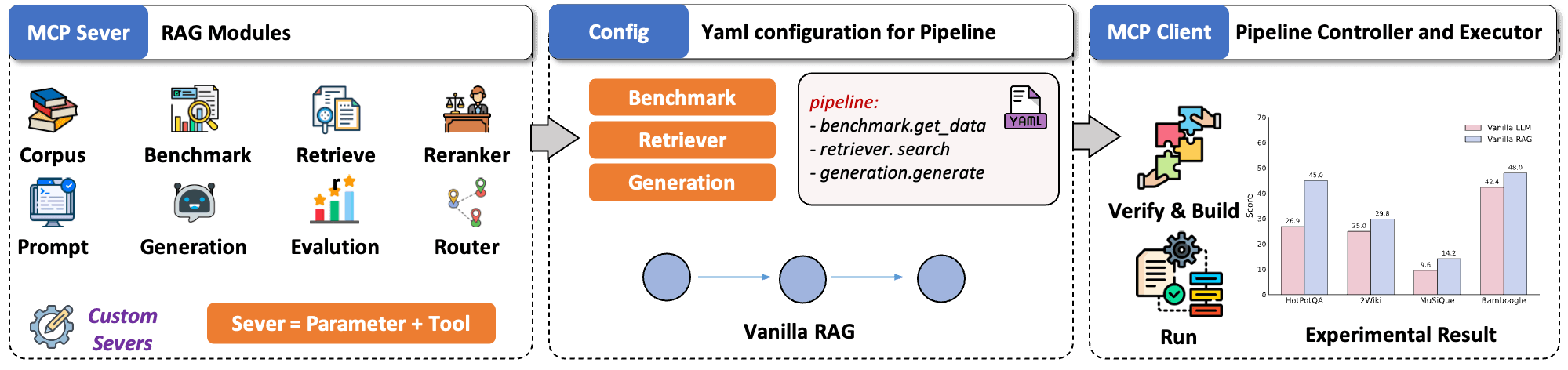
Why and How to Integrate Milvus into the UltraRAG Pipeline
In the UltraRAG v2 stack, the vector database plays a critical role in retrieval quality and system performance. Milvus, an open-source vector database, is a strong fit thanks to its scalability, efficient indexing, and seamless integration capabilities.
Once Milvus is integrated into an UltraRAG pipeline, you can build indexes and run queries using simple commands such as ultrarag build and ultrarag run. UltraRAG will automatically load your configuration and coordinate all modules needed to complete the task.
In this demo, we will walk through four objectives:
Integrate Milvus into an UltraRAG v2 project
Create a custom pipeline that uses Milvus for retrieval
Run the full pipeline to verify everything works end to end
Check the run results (Optional)
Dataset Setup
For this demo, we will use the Milvus FAQ dataset from the official Milvus repository. The dataset is provided in JSONL format.
{"id": "faq_0", "contents": "If you failed to pull the Milvus Docker image from Docker Hub, try adding other registry mirrors. Users from the Chinese mainland can add the URL https://registry.docker-cn.com to the registry-mirrors array in /etc.docker/daemon.json."}
{"id": "faq_1", "contents": "Docker is an efficient way to deploy Milvus, but not the only way. You can also deploy Milvus from source code. This requires Ubuntu (18.04 or higher) or CentOS (7 or higher). See Building Milvus from Source Code for more information."}
{"id": "faq_2", "contents": "Recall is affected mainly by index type and search parameters. For FLAT index, Milvus takes an exhaustive scan within a collection, with a 100% return. For IVF indexes, the nprobe parameter determines the scope of a search within the collection. Increasing nprobe increases the proportion of vectors searched and recall, but diminishes query performance."}
{"id": "faq_3", "contents": "Milvus does not support modification to configuration files during runtime. You must restart Milvus Docker for configuration file changes to take effect."}
{"id": "faq_4", "contents": "If Milvus is started using Docker Compose, run docker ps to observe how many Docker containers are running and check if Milvus services started correctly. For Milvus standalone, you should be able to observe at least three running Docker containers, one being the Milvus service and the other two being etcd management and storage service."}
{"id": "faq_5", "contents": "The time difference is usually due to the fact that the host machine does not use Coordinated Universal Time (UTC). The log files inside the Docker image use UTC by default. If your host machine does not use UTC, this issue may occur."}
{"id": "faq_6", "contents": "Milvus requires your CPU to support a SIMD instruction set: SSE4.2, AVX, AVX2, or AVX512. CPU must support at least one of these to ensure that Milvus operates normally."}
{"id": "faq_7", "contents": "Milvus requires your CPU to support a SIMD instruction set: SSE4.2, AVX, AVX2, or AVX512. CPU must support at least one of these to ensure that Milvus operates normally. An illegal instruction error returned during startup suggests that your CPU does not support any of the above four instruction sets."}
{"id": "faq_8", "contents": "Yes. You can install Milvus on Windows either by compiling from source code or from a binary package. See Run Milvus on Windows to learn how to install Milvus on Windows."}
{"id": "faq_9", "contents": "It is not recommended to install PyMilvus on Windows. But if you have to install PyMilvus on Windows but got an error, try installing it in a Conda environment. See Install Milvus SDK for more information about how to install PyMilvus in the Conda environment."}
Step 1: Deploy the Milvus Vector Database
Download the Deployment Files
wget https://github.com/Milvus-io/Milvus/releases/download/v2.5.12/Milvus-standalone-docker-compose.yml -O docker-compose.yml
Start the Milvus Service
docker-compose up -d
docker-compose ps -a
 Step 2: Clone the Project
Step 2: Clone the Project
git clone https://github.com/OpenBMB/UltraRAG.git
Step 3: Implement the Pipeline
Integrate the Milvus Vector Database
Note: Milvus is added as one of the supported vector database types in the retrieval module.
vim ultraRAG/UltraRAG/servers/retriever/src/retriever.py
import os
from urllib.parse import urlparse, urlunparse
from typing import Any, Dict, List, Optional
import aiohttp
import asyncio
import jsonlines
import numpy as np
import pandas as pd
from tqdm import tqdm
from flask import Flask, jsonify, request
from openai import AsyncOpenAI, OpenAIError
from fastmcp.exceptions import NotFoundError, ToolError, ValidationError
from ultrarag.server import UltraRAG_MCP_Server
app = UltraRAG_MCP_Server("retriever")
retriever_app = Flask(__name__)
class Retriever:
def __init__(self, mcp_inst: UltraRAG_MCP_Server):
mcp_inst.tool(
self.retriever_init,
output="retriever_path,corpus_path,index_path,faiss_use_gpu,infinity_kwargs,cuda_devices->None",
)
mcp_inst.tool(
self.retriever_init_openai,
output="corpus_path,openai_model,api_base,api_key->None",
)
mcp_inst.tool(
self.retriever_init_Milvus,
output="corpus_path,Milvus_host,Milvus_port,collection_name,embedding_dim->None",
)
mcp_inst.tool(
self.retriever_embed,
output="embedding_path,overwrite,use_alibaba_cloud,alibaba_api_key,alibaba_model,alibaba_endpoint->None",
)
mcp_inst.tool(
self.retriever_embed_openai,
output="embedding_path,overwrite->None",
)
mcp_inst.tool(
self.retriever_index,
output="embedding_path,index_path,overwrite,index_chunk_size->None",
)
mcp_inst.tool(
self.retriever_index_lancedb,
output="embedding_path,lancedb_path,table_name,overwrite->None",
)
# Note: retriever_index_Milvus has been removed
# Use setup_Milvus_collection.py for collection creation and indexing
mcp_inst.tool(
self.retriever_search,
output="q_ls,top_k,query_instruction,use_openai->ret_psg",
)
mcp_inst.tool(
self.retriever_search_lancedb,
output="q_ls,top_k,query_instruction,use_openai,lancedb_path,table_name,filter_expr->ret_psg",
)
mcp_inst.tool(
self.retriever_search_Milvus,
output="q_ls,top_k,query_instruction,use_openai->ret_psg",
)
mcp_inst.tool(
self.retriever_deploy_service,
output="retriever_url->None",
)
mcp_inst.tool(
self.retriever_deploy_search,
output="retriever_url,q_ls,top_k,query_instruction->ret_psg",
)
mcp_inst.tool(
self.retriever_exa_search,
output="q_ls,top_k->ret_psg",
)
mcp_inst.tool(
self.retriever_tavily_search,
output="q_ls,top_k->ret_psg",
)
def retriever_init(
self,
retriever_path: str,
corpus_path: str,
index_path: Optional[str] = None,
faiss_use_gpu: bool = False,
infinity_kwargs: Optional[Dict[str, Any]] = None,
cuda_devices: Optional[str] = None,
):
try:
import faiss
except ImportError:
err_msg = "faiss is not installed. Please install it with `conda install -c pytorch faiss-cpu` or `conda install -c pytorch faiss-gpu`."
app.logger.error(err_msg)
raise ImportError(err_msg)
try:
from infinity_emb.log_handler import LOG_LEVELS
from infinity_emb import AsyncEngineArray, EngineArgs
except ImportError:
err_msg = "infinity_emb is not installed. Please install it with `pip install infinity-emb`."
app.logger.error(err_msg)
raise ImportError(err_msg)
self.faiss_use_gpu = faiss_use_gpu
app.logger.setLevel(LOG_LEVELS["warning"])
if cuda_devices is not None:
assert isinstance(cuda_devices, str), "cuda_devices should be a string"
os.environ["CUDA_VISIBLE_DEVICES"] = cuda_devices
infinity_kwargs = infinity_kwargs or {}
self.model = AsyncEngineArray.from_args(
[EngineArgs(model_name_or_path=retriever_path, **infinity_kwargs)]
)[0]
self.contents = []
with jsonlines.open(corpus_path, mode="r") as reader:
self.contents = [item["contents"] for item in reader]
self.faiss_index = None
if index_path is not None and os.path.exists(index_path):
cpu_index = faiss.read_index(index_path)
if self.faiss_use_gpu:
co = faiss.GpuMultipleClonerOptions()
co.shard = True
co.useFloat16 = True
try:
self.faiss_index = faiss.index_cpu_to_all_gpus(cpu_index, co)
app.logger.info(f"Loaded index to GPU(s).")
except RuntimeError as e:
app.logger.error(
f"GPU index load failed: {e}. Falling back to CPU."
)
self.faiss_use_gpu = False
self.faiss_index = cpu_index
else:
self.faiss_index = cpu_index
app.logger.info("Loaded index on CPU.")
app.logger.info(f"Retriever index path has already been built")
else:
app.logger.warning(f"Cannot find path: {index_path}")
self.faiss_index = None
app.logger.info(f"Retriever initialized")
def retriever_init_openai(
self,
corpus_path: str,
openai_model: str,
api_base: str,
api_key: str,
):
if not openai_model:
raise ValueError("openai_model must be provided.")
if not api_base or not isinstance(api_base, str):
raise ValueError("api_base must be a non-empty string.")
if not api_key or not isinstance(api_key, str):
raise ValueError("api_key must be a non-empty string.")
self.contents = []
with jsonlines.open(corpus_path, mode="r") as reader:
self.contents = [item["contents"] for item in reader]
try:
self.openai_model = openai_model
self.client = AsyncOpenAI(base_url=api_base, api_key=api_key)
app.logger.info(
f"OpenAI client initialized with model '{openai_model}' and base '{api_base}'"
)
except OpenAIError as e:
app.logger.error(f"Failed to initialize OpenAI client: {e}")
def retriever_init_Milvus(
self,
corpus_path: str,
Milvus_host: str = "192.168.8.130",
Milvus_port: int = 19530,
collection_name: str = "ultrarag_collection_v3",
embedding_dim: int = 1024,
):
"""Initialize Milvus vector database connection.
Args:
corpus_path (str): Path to the corpus JSONL file (for reference only)
Milvus_host (str): Milvus server host
Milvus_port (int): Milvus server port
collection_name (str): Name of the existing collection to use
embedding_dim (int): Dimension of embeddings (for reference only)
Note:
This method assumes the collection already exists and is properly configured.
Use setup_Milvus_collection.py to create and configure collections.
"""
try:
from pyMilvus import connections, Collection, utility
except ImportError:
err_msg = "pyMilvus is not installed. Please install it with `pip install pyMilvus`."
app.logger.error(err_msg)
raise ImportError(err_msg)
# Initialize Alibaba Cloud client for embeddings
try:
from openai import AsyncOpenAI
except ImportError:
err_msg = "openai is not installed. Please install it with `pip install openai`."
app.logger.error(err_msg)
raise ImportError(err_msg)
# Set up Alibaba Cloud client for embeddings
self.alibaba_client = AsyncOpenAI(
base_url="https://dashscope.aliyuncs.com/compatible-mode/v1",
api_key="sk-xxxxxx"
)
self.alibaba_model = "text-embedding-v3"
# Load corpus data (for reference, not used in search)
self.contents = []
with jsonlines.open(corpus_path, mode="r") as reader:
self.contents = [item["contents"] for item in reader]
# Connect to Milvus
try:
connections.connect(
alias="default",
host=Milvus_host,
port=Milvus_port
)
app.logger.info(f"Connected to Milvus at {Milvus_host}:{Milvus_port}")
except Exception as e:
app.logger.error(f"Failed to connect to Milvus: {e}")
raise ConnectionError(f"Failed to connect to Milvus: {e}")
# Store Milvus configuration
self.Milvus_host = Milvus_host
self.Milvus_port = Milvus_port
self.collection_name = collection_name
self.embedding_dim = embedding_dim
# Connect to existing collection (must exist and be loaded)
if not utility.has_collection(collection_name):
raise ValueError(f"Collection '{collection_name}' does not exist. Please create it first using setup_Milvus_collection.py")
self.Milvus_collection = Collection(collection_name)
# Verify collection is loaded
load_state = utility.load_state(collection_name)
if load_state != "Loaded":
app.logger.warning(f"Collection '{collection_name}' is not loaded (state: {load_state}). Attempting to load...")
try:
self.Milvus_collection.load()
utility.wait_for_loading_complete(collection_name=collection_name, timeout=60)
app.logger.info(f"Successfully loaded collection '{collection_name}'")
except Exception as e:
raise RuntimeError(f"Failed to load collection '{collection_name}': {e}")
# Verify collection has data and indexes
entity_count = self.Milvus_collection.num_entities
if entity_count == 0:
app.logger.warning(f"Collection '{collection_name}' is empty")
else:
app.logger.info(f"Connected to collection '{collection_name}' with {entity_count} entities")
app.logger.info("Milvus retriever initialized successfully")
async def retriever_embed(
self,
embedding_path: Optional[str] = None,
overwrite: bool = False,
use_alibaba_cloud: bool = False,
alibaba_api_key: Optional[str] = None,
alibaba_model: str = "text-embedding-v3",
alibaba_endpoint: Optional[str] = None,
):
if embedding_path is not None:
if not embedding_path.endswith(".npy"):
err_msg = f"Embedding save path must end with .npy, now the path is {embedding_path}"
app.logger.error(err_msg)
raise ValidationError(err_msg)
output_dir = os.path.dirname(embedding_path)
else:
current_file = os.path.abspath(__file__)
project_root = os.path.dirname(os.path.dirname(current_file))
output_dir = os.path.join(project_root, "output", "embedding")
embedding_path = os.path.join(output_dir, "embedding.npy")
if not overwrite and os.path.exists(embedding_path):
app.logger.info("embedding already exists, skipping")
return
os.makedirs(output_dir, exist_ok=True)
if use_alibaba_cloud:
# Use Alibaba Cloud API for embeddings
if not alibaba_api_key or not alibaba_endpoint:
raise ValueError("Alibaba Cloud API key and endpoint must be provided")
client = AsyncOpenAI(base_url=alibaba_endpoint, api_key=alibaba_api_key)
async def alibaba_embed(texts):
embeddings = []
batch_size = 100 # Process in batches to avoid rate limits
for i in range(0, len(texts), batch_size):
batch = texts[i:i+batch_size]
try:
response = await client.embeddings.create(
input=batch, model=alibaba_model
)
batch_embeddings = [item.embedding for item in response.data]
embeddings.extend(batch_embeddings)
app.logger.info(f"Processed batch {i//batch_size + 1}/{(len(texts)-1)//batch_size + 1}")
except Exception as e:
app.logger.error(f"Error in Alibaba Cloud embedding batch {i//batch_size + 1}: {e}")
raise
return embeddings
embeddings = await alibaba_embed(self.contents)
app.logger.info("Alibaba Cloud embedding completed")
else:
# Use local model for embeddings
async with self.model:
embeddings, usage = await self.model.embed(sentences=self.contents)
embeddings = np.array(embeddings, dtype=np.float16)
np.save(embedding_path, embeddings)
app.logger.info("embedding success")
async def retriever_embed_openai(
self,
embedding_path: Optional[str] = None,
overwrite: bool = False,
):
if embedding_path is not None:
if not embedding_path.endswith(".npy"):
err_msg = f"Embedding save path must end with .npy, now the path is {embedding_path}"
app.logger.error(err_msg)
raise ValidationError(err_msg)
output_dir = os.path.dirname(embedding_path)
else:
current_file = os.path.abspath(__file__)
project_root = os.path.dirname(os.path.dirname(current_file))
output_dir = os.path.join(project_root, "output", "embedding")
embedding_path = os.path.join(output_dir, "embedding.npy")
if not overwrite and os.path.exists(embedding_path):
app.logger.info("embedding already exists, skipping")
os.makedirs(output_dir, exist_ok=True)
async def openai_embed(texts):
embeddings = []
for text in texts:
response = await self.client.embeddings.create(
input=text, model=self.openai_model
)
embeddings.append(response.data[0].embedding)
return embeddings
embeddings = await openai_embed(self.contents)
embeddings = np.array(embeddings, dtype=np.float16)
np.save(embedding_path, embeddings)
app.logger.info("embedding success")
def retriever_index(
self,
embedding_path: str,
index_path: Optional[str] = None,
overwrite: bool = False,
index_chunk_size: int = 50000,
):
"""
Build a Faiss index from an embedding matrix.
Args:
embedding_path (str): .npy file of shape (N, dim), dtype float32.
index_path (str, optional): where to save .index file.
overwrite (bool): overwrite existing index.
index_chunk_size (int): batch size for add_with_ids.
"""
try:
import faiss
except ImportError:
err_msg = "faiss is not installed. Please install it with `conda install -c pytorch faiss-cpu` or `conda install -c pytorch faiss-gpu`."
app.logger.error(err_msg)
raise ImportError(err_msg)
if not os.path.exists(embedding_path):
app.logger.error(f"Embedding file not found: {embedding_path}")
NotFoundError(f"Embedding file not found: {embedding_path}")
if index_path is not None:
if not index_path.endswith(".index"):
app.logger.error(
f"Parameter index_path must end with .index now is {index_path}"
)
ValidationError(
f"Parameter index_path must end with .index now is {index_path}"
)
output_dir = os.path.dirname(index_path)
else:
current_file = os.path.abspath(__file__)
project_root = os.path.dirname(os.path.dirname(current_file))
output_dir = os.path.join(project_root, "output", "index")
index_path = os.path.join(output_dir, "index.index")
if not overwrite and os.path.exists(index_path):
app.logger.info("Index already exists, skipping")
os.makedirs(output_dir, exist_ok=True)
embedding = np.load(embedding_path)
dim = embedding.shape[1]
vec_ids = np.arange(embedding.shape[0]).astype(np.int64)
# with cpu
cpu_flat = faiss.IndexFlatIP(dim)
cpu_index = faiss.IndexIDMap2(cpu_flat)
# chunk to write
total = embedding.shape[0]
for start in range(0, total, index_chunk_size):
end = min(start + index_chunk_size, total)
cpu_index.add_with_ids(embedding[start:end], vec_ids[start:end])
# with gpu
if self.faiss_use_gpu:
co = faiss.GpuMultipleClonerOptions()
co.shard = True
co.useFloat16 = True
try:
gpu_index = faiss.index_cpu_to_all_gpus(cpu_index, co)
index = gpu_index
app.logger.info("Using GPU for indexing with sharding")
except RuntimeError as e:
app.logger.warning(f"GPU indexing failed ({e}); fall back to CPU")
self.faiss_use_gpu = False
index = cpu_index
else:
index = cpu_index
# save
faiss.write_index(cpu_index, index_path)
if self.faiss_index is None:
self.faiss_index = index
app.logger.info("Indexing success")
def retriever_index_lancedb(
self,
embedding_path: str,
lancedb_path: str,
table_name: str,
overwrite: bool = False,
):
"""
Build a Faiss index from an embedding matrix.
Args:
embedding_path (str): .npy file of shape (N, dim), dtype float32.
lancedb_path (str): directory path to store LanceDB tables.
table_name (str): the name of the LanceDB table.
overwrite (bool): overwrite existing index.
"""
try:
import lancedb
except ImportError:
err_msg = "lancedb is not installed. Please install it with `pip install lancedb`."
app.logger.error(err_msg)
raise ImportError(err_msg)
if not os.path.exists(embedding_path):
app.logger.error(f"Embedding file not found: {embedding_path}")
NotFoundError(f"Embedding file not found: {embedding_path}")
if lancedb_path is None:
current_file = os.path.abspath(__file__)
project_root = os.path.dirname(os.path.dirname(current_file))
lancedb_path = os.path.join(project_root, "output", "lancedb")
os.makedirs(lancedb_path, exist_ok=True)
db = lancedb.connect(lancedb_path)
if table_name in db.table_names() and not overwrite:
info_msg = f"LanceDB table '{table_name}' already exists, skipping"
app.logger.info(info_msg)
return {"status": info_msg}
elif table_name in db.table_names() and overwrite:
import shutil
shutil.rmtree(os.path.join(lancedb_path, table_name))
app.logger.info(f"Overwriting LanceDB table '{table_name}'")
embedding = np.load(embedding_path)
ids = [str(i) for i in range(len(embedding))]
data = [{"id": i, "vector": v} for i, v in zip(ids, embedding)]
df = pd.DataFrame(data)
db.create_table(table_name, data=df)
app.logger.info("LanceDB indexing success")
# Note: retriever_index_Milvus method has been removed
# Collection creation and indexing is now handled by setup_Milvus_collection.py
# This simplifies the retriever logic and separates concerns
async def retriever_search(
self,
query_list: List[str],
top_k: int = 5,
query_instruction: str = "",
use_openai: bool = False,
) -> Dict[str, List[List[str]]]:
if isinstance(query_list, str):
query_list = [query_list]
queries = [f"{query_instruction}{query}" for query in query_list]
if use_openai:
async def openai_embed(texts):
embeddings = []
for text in texts:
response = await self.client.embeddings.create(
input=text, model=self.openai_model
)
embeddings.append(response.data[0].embedding)
return embeddings
query_embedding = await openai_embed(queries)
else:
async with self.model:
query_embedding, usage = await self.model.embed(sentences=queries)
query_embedding = np.array(query_embedding, dtype=np.float16)
app.logger.info("query embedding finish")
scores, ids = self.faiss_index.search(query_embedding, top_k)
rets = []
for i, query in enumerate(query_list):
cur_ret = []
for _, id in enumerate(ids[i]):
cur_ret.append(self.contents[id])
rets.append(cur_ret)
app.logger.debug(f"ret_psg: {rets}")
return {"ret_psg": rets}
async def retriever_search_Milvus(
self,
query_list: List[str],
top_k: int = 5,
query_instruction: str = "",
use_openai: bool = False,
) -> Dict[str, List[List[str]]]:
"""
Search in Milvus vector database.
Args:
query_list (List[str]): List of query strings
top_k (int): Number of top results to return
query_instruction (str): Instruction to prepend to queries
use_openai (bool): Whether to use OpenAI for embedding
Returns:
Dict[str, List[List[str]]]: Search results
"""
try:
from pyMilvus import connections, Collection
except ImportError:
err_msg = "pyMilvus is not installed. Please install it with `pip install pyMilvus`."
app.logger.error(err_msg)
raise ImportError(err_msg)
if isinstance(query_list, str):
query_list = [query_list]
queries = [f"{query_instruction}{query}" for query in query_list]
# Generate query embeddings
if use_openai:
async def openai_embed(texts):
embeddings = []
for text in texts:
response = await self.client.embeddings.create(
input=text, model=self.openai_model
)
embeddings.append(response.data[0].embedding)
return embeddings
query_embedding = await openai_embed(queries)
else:
# Use Alibaba Cloud API for embeddings
async def alibaba_embed(texts):
embeddings = []
for text in texts:
response = await self.alibaba_client.embeddings.create(
input=text, model=self.alibaba_model
)
embeddings.append(response.data[0].embedding)
return embeddings
query_embedding = await alibaba_embed(queries)
query_embedding = np.array(query_embedding, dtype=np.float32)
app.logger.info("Query embedding finished")
# Ensure collection is loaded before search
try:
if not self.Milvus_collection.has_index():
app.logger.warning("Collection has no index, search may be slow")
# Always load collection before search to ensure it's available
app.logger.debug("Loading collection for search...")
self.Milvus_collection.load()
app.logger.debug("Collection loaded successfully")
except Exception as load_error:
app.logger.error(f"Failed to load collection: {load_error}")
return {"ret_psg": [[]] * len(query_list)}
# Search in Milvus
search_params = {
"metric_type": "IP",
"params": {"nprobe": 10}
}
rets = []
for i, query_vec in enumerate(query_embedding):
try:
# Perform search with proper error handling
results = self.Milvus_collection.search(
data=[query_vec.tolist()],
anns_field="embedding",
param=search_params,
limit=top_k,
output_fields=["text"],
expr=None # Explicitly set no filter expression
)
# Extract results with null checks
cur_ret = []
for hit in results[0]:
text_content = hit.entity.get("text")
if text_content is not None:
cur_ret.append(text_content)
else:
app.logger.warning(f"Found null text content in search result")
rets.append(cur_ret)
except Exception as e:
app.logger.error(f"Milvus search failed for query {i}: {e}")
# Return empty result for failed query
rets.append([])
app.logger.debug(f"ret_psg: {rets}")
return {"ret_psg": rets}
async def retriever_search_lancedb(
self,
query_list: List[str],
top_k: Optional[int] | None = None,
query_instruction: str = "",
use_openai: bool = False,
lancedb_path: str = "",
table_name: str = "",
filter_expr: Optional[str] = None,
) -> Dict[str, List[List[str]]]:
try:
import lancedb
except ImportError:
err_msg = "lancedb is not installed. Please install it with `pip install lancedb`."
app.logger.error(err_msg)
raise ImportError(err_msg)
if isinstance(query_list, str):
query_list = [query_list]
queries = [f"{query_instruction}{query}" for query in query_list]
if use_openai:
async def openai_embed(texts):
embeddings = []
for text in texts:
response = await self.client.embeddings.create(
input=text, model=self.openai_model
)
embeddings.append(response.data[0].embedding)
return embeddings
query_embedding = await openai_embed(queries)
else:
async with self.model:
query_embedding, usage = await self.model.embed(sentences=queries)
query_embedding = np.array(query_embedding, dtype=np.float16)
app.logger.info("query embedding finish")
rets = []
if not lancedb_path:
NotFoundError(f"`lancedb_path` must be provided.")
db = lancedb.connect(lancedb_path)
self.lancedb_table = db.open_table(table_name)
for i, query_vec in enumerate(query_embedding):
q = self.lancedb_table.search(query_vec).limit(top_k)
if filter_expr:
q = q.where(filter_expr)
df = q.to_df()
cur_ret = []
for id_str in df["id"]:
id_int = int(id_str)
cur_ret.append(self.contents[id_int])
rets.append(cur_ret)
app.logger.debug(f"ret_psg: {rets}")
return {"ret_psg": rets}
async def retriever_deploy_service(
self,
retriever_url: str,
):
# Ensure URL is valid, adding "http://" prefix if necessary
retriever_url = retriever_url.strip()
if not retriever_url.startswith("http://") and not retriever_url.startswith(
"https://"
):
retriever_url = f"http://{retriever_url}"
url_obj = urlparse(retriever_url)
retriever_host = url_obj.hostname
retriever_port = (
url_obj.port if url_obj.port else 8080
) # Default port if none provided
@retriever_app.route("/search", methods=["POST"])
async def deploy_retrieval_model():
data = request.get_json()
query_list = data["query_list"]
top_k = data["top_k"]
async with self.model:
query_embedding, _ = await self.model.embed(sentences=query_list)
query_embedding = np.array(query_embedding, dtype=np.float16)
_, ids = self.faiss_index.search(query_embedding, top_k)
rets = []
for i, _ in enumerate(query_list):
cur_ret = []
for _, id in enumerate(ids[i]):
cur_ret.append(self.contents[id])
rets.append(cur_ret)
return jsonify({"ret_psg": rets})
retriever_app.run(host=retriever_host, port=retriever_port)
app.logger.info(f"employ embedding server at {retriever_url}")
async def retriever_deploy_search(
self,
retriever_url: str,
query_list: List[str],
top_k: Optional[int] | None = None,
query_instruction: str = "",
):
# Validate the URL format
url = retriever_url.strip()
if not url.startswith("http://") and not url.startswith("https://"):
url = f"http://{url}"
url_obj = urlparse(url)
api_url = urlunparse(url_obj._replace(path="/search"))
app.logger.info(f"Calling url: {api_url}")
if isinstance(query_list, str):
query_list = [query_list]
query_list = [f"{query_instruction}{query}" for query in query_list]
payload = {"query_list": query_list}
if top_k is not None:
payload["top_k"] = top_k
async with aiohttp.ClientSession() as session:
async with session.post(
api_url,
json=payload,
) as response:
if response.status == 200:
response_data = await response.json()
app.logger.debug(
f"status_code: {response.status}, response data: {response_data}"
)
return response_data
else:
err_msg = (
f"Failed to call {retriever_url} with code {response.status}"
)
app.logger.error(err_msg)
raise ToolError(err_msg)
async def retriever_exa_search(
self,
query_list: List[str],
top_k: Optional[int] | None = None,
) -> dict[str, List[List[str]]]:
try:
from exa_py import AsyncExa
from exa_py.api import Result
except ImportError:
err_msg = (
"exa_py is not installed. Please install it with `pip install exa_py`."
)
app.logger.error(err_msg)
raise ImportError(err_msg)
exa_api_key = os.environ.get("EXA_API_KEY", "")
exa = AsyncExa(api_key=exa_api_key if exa_api_key else "EMPTY")
sem = asyncio.Semaphore(16)
async def call_with_retry(
idx: int, q: str, retries: int = 3, delay: float = 1.0
):
async with sem:
for attempt in range(retries):
try:
resp = await exa.search_and_contents(
q,
num_results=top_k,
text=True,
)
results: List[Result] = getattr(resp, "results", []) or []
psg_ls: List[str] = [(r.text or "") for r in results]
return idx, psg_ls
except Exception as e:
status = getattr(
getattr(e, "response", None), "status_code", None
)
if status == 401 or "401" in str(e):
raise RuntimeError(
"Unauthorized (401): Access denied by Exa API. "
"Invalid or missing EXA_API_KEY."
) from e
app.logger.warning(
f"[Retry {attempt+1}] EXA failed (idx={idx}): {e}"
)
await asyncio.sleep(delay)
return idx, []
tasks = [
asyncio.create_task(call_with_retry(i, q)) for i, q in enumerate(query_list)
]
ret: List[List[str]] = [None] * len(query_list)
iterator = tqdm(
asyncio.as_completed(tasks), total=len(tasks), desc="EXA Searching: "
)
for fut in iterator:
idx, psg_ls = await fut
ret[idx] = psg_ls
return {"ret_psg": ret}
async def retriever_tavily_search(
self,
query_list: List[str],
top_k: Optional[int] | None = None,
) -> dict[str, List[List[str]]]:
try:
from tavily import (
AsyncTavilyClient,
BadRequestError,
UsageLimitExceededError,
InvalidAPIKeyError,
MissingAPIKeyError,
)
except ImportError:
err_msg = "tavily is not installed. Please install it with `pip install tavily-python`."
app.logger.error(err_msg)
raise ImportError(err_msg)
tavily_api_key = os.environ.get("TAVILY_API_KEY", "")
if not tavily_api_key:
raise MissingAPIKeyError(
"TAVILY_API_KEY environment variable is not set. Please set it to use Tavily."
)
tavily = AsyncTavilyClient(api_key=tavily_api_key)
sem = asyncio.Semaphore(16)
async def call_with_retry(
idx: int, q: str, retries: int = 3, delay: float = 1.0
):
async with sem:
for attempt in range(retries):
try:
resp = await tavily.search(
query=q,
max_results=top_k,
)
results: List[Dict[str, Any]] = resp["results"]
psg_ls: List[str] = [(r["content"] or "") for r in results]
return idx, psg_ls
except UsageLimitExceededError as e:
app.logger.error(f"Usage limit exceeded: {e}")
raise ToolError(f"Usage limit exceeded: {e}") from e
except InvalidAPIKeyError as e:
app.logger.error(f"Invalid API key: {e}")
raise ToolError(f"Invalid API key: {e}") from e
except (BadRequestError, Exception) as e:
app.logger.warning(
f"[Retry {attempt+1}] Tavily failed (idx={idx}): {e}"
)
await asyncio.sleep(delay)
return idx, []
tasks = [
asyncio.create_task(call_with_retry(i, q)) for i, q in enumerate(query_list)
]
ret: List[List[str]] = [None] * len(query_list)
iterator = tqdm(
asyncio.as_completed(tasks), total=len(tasks), desc="Tavily Searching: "
)
for fut in iterator:
idx, psg_ls = await fut
ret[idx] = psg_ls
return {"ret_psg": ret}
if __name__ == "__main__":
Retriever(app)
app.run(transport="stdio")
Define the Parameter Configuration File
Note: This file specifies all parameter settings used in the pipeline.
vim parameter.yaml
# servers/retriever/parameter.yaml
retriever_path: openbmb/MiniCPM-Embedding-Light
corpus_path: UltraRAG/data/Milvus_faq_corpus.jsonl
embedding_path: embedding/embedding.npy
index_path: index/index.index
# infinify_emb config
infinity_kwargs:
bettertransformer: false
pooling_method: auto
device: cuda
batch_size: 1024
cuda_devices: "0,1"
query_instruction: "Query: "
faiss_use_gpu: True
top_k: 5
overwrite: false
retriever_url: http://localhost:8080
index_chunk_size: 50000
# OpenAI API configuration (if used)
use_openai: false
openai_model: "embedding"
api_base: ""
api_key: ""
# Alibaba Cloud API configuration (alternative to local embedding)
use_alibaba_cloud: true
alibaba_api_key: "sk-xxxxxxx" # Your Alibaba Cloud API key
alibaba_model: "embedding" # Alibaba Cloud embedding model
alibaba_endpoint: "https://dashscope.aliyuncs.com/compatible-mode/v1" # Alibaba Cloud endpoint
# LanceDB configuration (if used)
lancedb_path: "lancedb/"
table_name: "vector_index"
filter_expr: null
# Milvus configuration (if used)
use_Milvus: true
Milvus_host: "192.168.8.130"
Milvus_port: 19530
collection_name: "ultrarag_collection_v3"
embedding_dim: 1024
Define the Server Configuration File
Note: This configuration includes integration with Alibaba Cloud APIs.
vim rag_Milvus_faq_server.yaml
benchmark:
parameter: /root/ultraRAG/UltraRAG/servers/benchmark/parameter.yaml
path: /root/ultraRAG/UltraRAG/servers/benchmark/src/benchmark.py
tools:
get_data:
input:
benchmark: $benchmark
output:
- q_ls
- gt_ls
custom:
parameter: /root/ultraRAG/UltraRAG/servers/custom/parameter.yaml
path: /root/ultraRAG/UltraRAG/servers/custom/src/custom.py
tools:
output_extract_from_boxed:
input:
ans_ls: ans_ls
output:
- pred_ls
evaluation:
parameter: /root/ultraRAG/UltraRAG/servers/evaluation/parameter.yaml
path: /root/ultraRAG/UltraRAG/servers/evaluation/src/evaluation.py
tools:
evaluate:
input:
gt_ls: gt_ls
metrics: $metrics
pred_ls: pred_ls
save_path: $save_path
output:
- eval_res
generation:
parameter: /root/ultraRAG/UltraRAG/servers/generation/parameter.yaml
path: /root/ultraRAG/UltraRAG/servers/generation/src/generation.py
tools:
generate:
input:
base_url: $base_url
model_name: $model_name
prompt_ls: prompt_ls
sampling_params: $sampling_params
api_key: $api_key
output:
- ans_ls
prompt:
parameter: /root/ultraRAG/UltraRAG/servers/prompt/parameter.yaml
path: /root/ultraRAG/UltraRAG/servers/prompt/src/prompt.py
prompts:
qa_rag_boxed:
input:
q_ls: q_ls
ret_psg: ret_psg
template: $template
output:
- prompt_ls
retriever:
parameter: /root/ultraRAG/UltraRAG/servers/retriever/parameter.yaml
path: /root/ultraRAG/UltraRAG/servers/retriever/src/retriever.py
tools:
retriever_init_Milvus:
input:
collection_name: $collection_name
corpus_path: $corpus_path
embedding_dim: $embedding_dim
Milvus_host: $Milvus_host
Milvus_port: $Milvus_port
retriever_search_Milvus:
input:
query_instruction: $query_instruction
query_list: q_ls
top_k: $top_k
use_openai: $use_openai
output:
- ret_psg
Define the Build Index
Note: Convert the document corpus into vector embeddings and store them in Milvus. Configure the key indexing parameters needed for this process.
vim Milvus_index_parameter.yaml
retriever:
alibaba_api_key: sk-xxxxxxx
alibaba_endpoint: https://dashscope.aliyuncs.com/compatible-mode/v1
alibaba_model: text-embedding-v3
collection_name: ultrarag_collection_v3
corpus_path: data/corpus_example.jsonl
embedding_dim: 1024
embedding_path: embedding/embedding.npy
Milvus_host: 192.168.8.130
Milvus_port: 19530
overwrite: false
use_alibaba_cloud: true
vim mivus_index.yaml
# Milvus Index Building Configuration
# Build vector index using Milvus database
# Note: This configuration is now deprecated. Use setup_Milvus_collection.py instead.
# MCP Server
servers:
retriever: servers/retriever
# Parameter Configuration
parameter_config: examples/parameter/Milvus_index_parameter.yaml
# MCP Client Pipeline
# Updated pipeline for new architecture
pipeline:
- retriever.retriever_init_Milvus # Connect to existing Milvus collection
- retriever.retriever_embed # Generate embeddings (if needed)
# Note: Index building is now handled by setup_Milvus_collection.py
# The collection ultrarag_collection_v3 should already exist with proper indexing
Run RAG
Runs the full RAG workflow including retrieval, answer generation, and evaluation. Configure the key indexing parameters needed for this process.
vim rag_Milvus_faq_server
benchmark:
parameter: /root/ultraRAG/UltraRAG/servers/benchmark/parameter.yaml
path: /root/ultraRAG/UltraRAG/servers/benchmark/src/benchmark.py
tools:
get_data:
input:
benchmark: $benchmark
output:
- q_ls
- gt_ls
custom:
parameter: /root/ultraRAG/UltraRAG/servers/custom/parameter.yaml
path: /root/ultraRAG/UltraRAG/servers/custom/src/custom.py
tools:
output_extract_from_boxed:
input:
ans_ls: ans_ls
output:
- pred_ls
evaluation:
parameter: /root/ultraRAG/UltraRAG/servers/evaluation/parameter.yaml
path: /root/ultraRAG/UltraRAG/servers/evaluation/src/evaluation.py
tools:
evaluate:
input:
gt_ls: gt_ls
metrics: $metrics
pred_ls: pred_ls
save_path: $save_path
output:
- eval_res
generation:
parameter: /root/ultraRAG/UltraRAG/servers/generation/parameter.yaml
path: /root/ultraRAG/UltraRAG/servers/generation/src/generation.py
tools:
generate:
input:
base_url: $base_url
model_name: $model_name
prompt_ls: prompt_ls
sampling_params: $sampling_params
api_key: $api_key
output:
- ans_ls
prompt:
parameter: /root/ultraRAG/UltraRAG/servers/prompt/parameter.yaml
path: /root/ultraRAG/UltraRAG/servers/prompt/src/prompt.py
prompts:
qa_rag_boxed:
input:
q_ls: q_ls
ret_psg: ret_psg
template: $template
output:
- prompt_ls
retriever:
parameter: /root/ultraRAG/UltraRAG/servers/retriever/parameter.yaml
path: /root/ultraRAG/UltraRAG/servers/retriever/src/retriever.py
tools:
retriever_init_Milvus:
input:
collection_name: $collection_name
corpus_path: $corpus_path
embedding_dim: $embedding_dim
Milvus_host: $Milvus_host
Milvus_port: $Milvus_port
retriever_search_Milvus:
input:
query_instruction: $query_instruction
query_list: q_ls
top_k: $top_k
use_openai: $use_openai
output:
- ret_psg
vim rag_Milvus_faq.yaml
# Milvus RAG FAQ Demo
# Complete RAG pipeline using Milvus vector database with FAQ dataset
# MCP Server Configuration
servers:
benchmark: UltraRAG/servers/benchmark
retriever: UltraRAG/servers/retriever
prompt: UltraRAG/servers/prompt
generation: UltraRAG/servers/generation
evaluation: UltraRAG/servers/evaluation
custom: UltraRAG/servers/custom
# Parameter Configuration
parameter_config: examples/parameter/rag_Milvus_faq_parameter.yaml
# MCP Client Pipeline
# Sequential execution: data -> init -> search -> prompt -> generate -> extract -> evaluate
pipeline:
- benchmark.get_data
- retriever.retriever_init_Milvus
- retriever.retriever_search_Milvus
- prompt.qa_rag_boxed
- generation.generate
- custom.output_extract_from_boxed
- evaluation.evaluate
Run the Build Index
Note: After a successful run, the system generates the vector embeddings and index files. The RAG pipeline can then use these directly to perform retrieval.
ultrarag build examples/Milvus_index.yaml
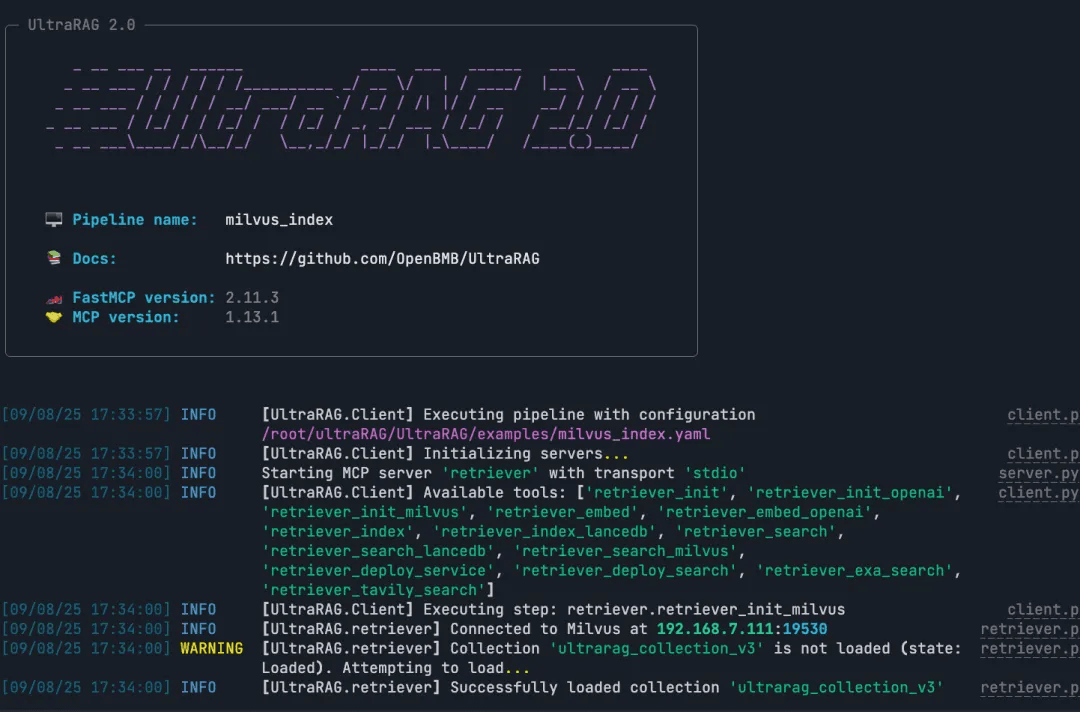
ultrarag run examples/Milvus_index.yaml
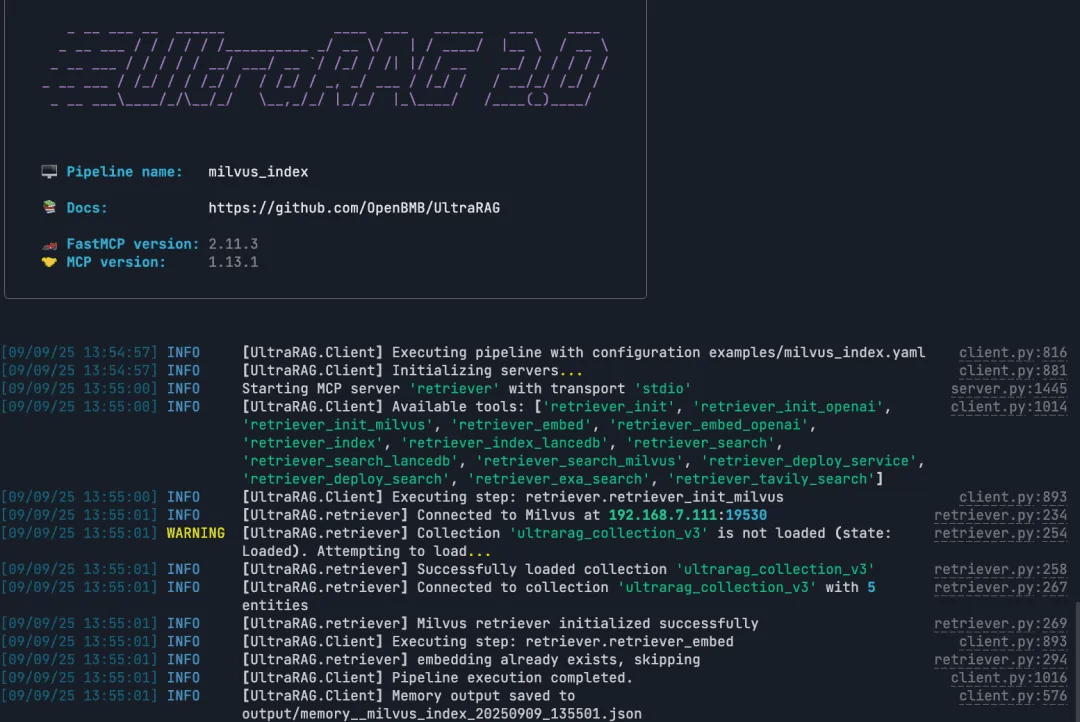
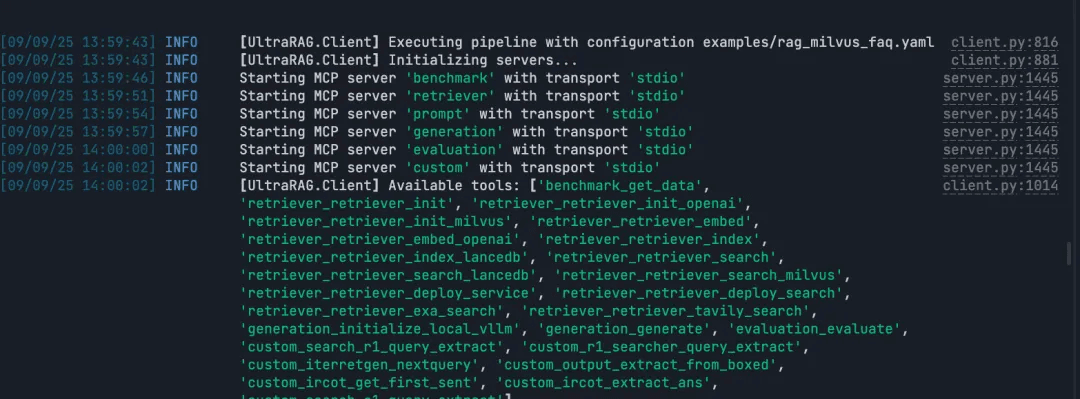
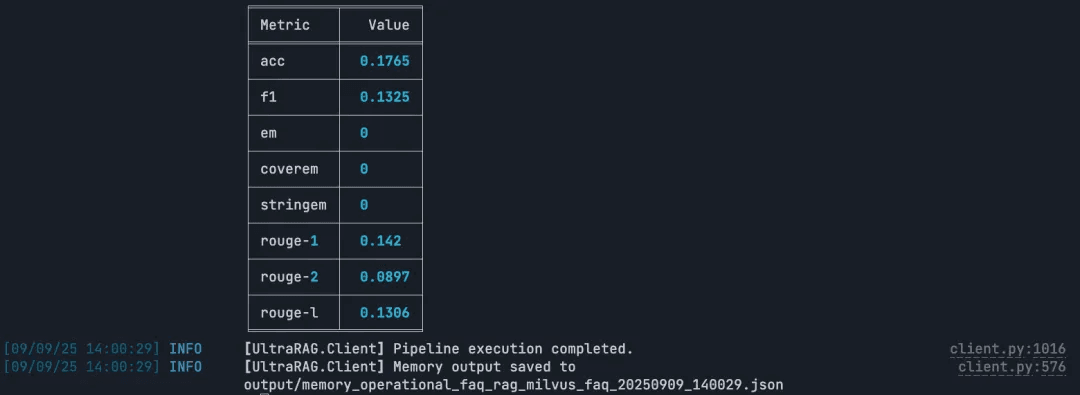
Run the RAG Query
Note: Build and execute the full RAG pipeline end to end.
ultrarag build examples/rag_Milvus.yaml
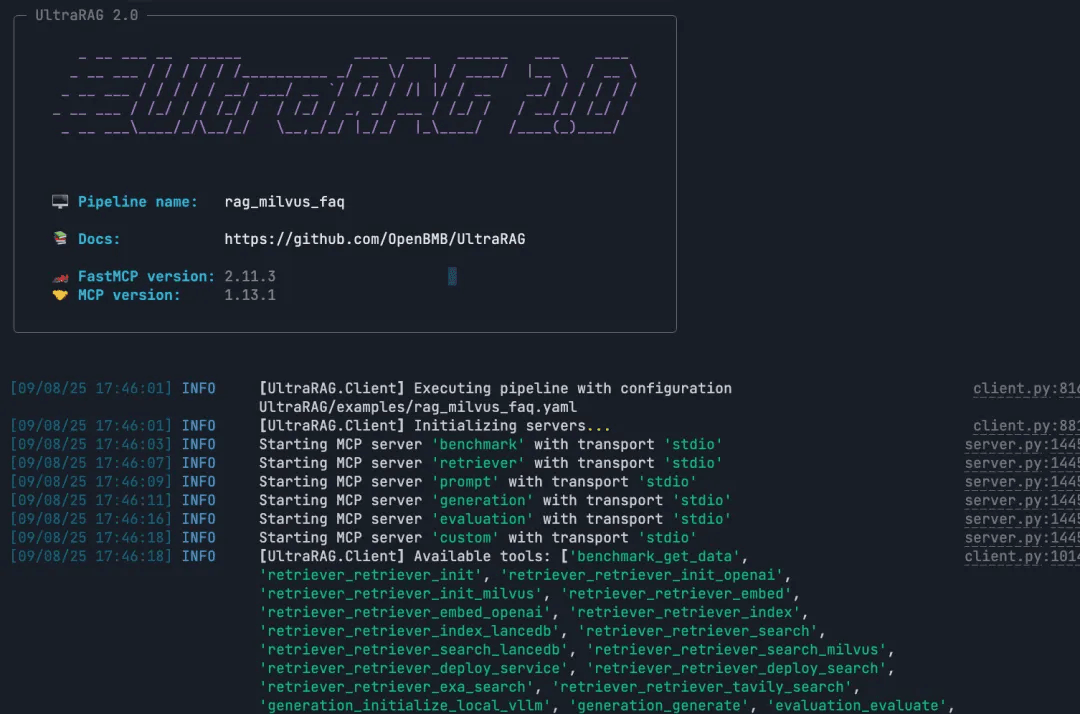
ultrarag run examples/rag_Milvus.yaml
Conclusion
Building a traditional RAG pipeline often means writing hundreds or even thousands of lines of code. UltraRAG v2 takes a very different approach. With its MCP-based modular design, declarative YAML configuration, and lightweight orchestration model, you can build the same end-to-end pipeline in just a few dozen lines. Also, YAML is used to describe the steps of the workflow, you don’t have to actually write real code. This make it much easier to bring enterprise-level RAG into real-world applications.
Integrating Milvus into this workflow adds another advantage: a high-performance, production-ready vector database that’s designed specifically for scalable semantic retrieval. Together, Milvus and UltraRAG v2 make it far easier to prototype quickly, iterate confidently, and deploy RAG systems that can handle real workloads.
Ready to simplify your RAG development? Try UltraRAG v2 with Milvus. Explore the example pipeline, run it yourself, and build a complete RAG workflow with just a few lines of configuration.
If you have any questions, join our Slack channel or book a 20-minute Milvus Office Hours session to discuss your use case.
- UltraRAG v2 Architecture at a Glance
- Why and How to Integrate Milvus into the UltraRAG Pipeline
- Dataset Setup
- Step 1: Deploy the Milvus Vector Database
- Step 2: Clone the Project
- Step 3: Implement the Pipeline
- Conclusion
On This Page
Try Managed Milvus for Free
Zilliz Cloud is hassle-free, powered by Milvus and 10x faster.
Get StartedLike the article? Spread the word



